
Int J Chem Res, Vol 6, Issue 4, 5-13Research Article
MOLECULAR DOCKING STUDIES OF MONOMERIC WILDTYPE AND MUTANT (H81A, H49R) SOD1 WITH EDARAVONE AND RILUZOLE
RADHIKA NAGRE1*, VAEESHNAVI BUWA2
1Department of Biochemistry, Osmania University, India, 2Bioinformatics Associate, BioNome, Bangalore, India
Email: [email protected]
Received: 10 Jun 2022 Revised and Accepted: 26 Jul 2022
ABSTRACT
Objective: Amyotrophic lateral sclerosis (ALS) is a fatal motor neuron disorder that causes progressive loss of the upper and lower motor neurons in the brain and spinal cord. SOD1 was the first gene linked to ALS, in which more than 150 mutations throughout the sequence of the protein have been found to be associated with ALS.
Methods: The drugs that can interact and inhibit the misfolding or revert the misconformation of the protein can be useful in the treatment of ALS. Monomer apo SOD1-WT and MBR SOD1 mutants (H81A, H49R) were docked with the only two FDA-approved drugs for ALS that are edaravone and riluzole to assess if the ALS patients carrying these particular protein aggregates will derive any therapeutic efficacy.
Results: The drugs were found to interact with both wild-type and mutant SOD1 at different positions and the type of interaction, degree of interaction and their binding energies were determined. SOD1-WT has hydrophobic interactions at V103, H110, and R115 and hydrogen bond at H110 with edaravone. SOD1-WT has hydrophobic interactions at T54, F64 and hydrogen bond at D52, T58 with riluzole. Both SOD1 mutants has a hydrogen bond at its H46 residue for both drugs. SOD1-H49R mutant has hydrophobic interactions at F45 with riluzole. SOD1-H81A mutant has hydrophobic interactions at F45 with edaravone.
Conclusion: Both the mutants L42A residue has a hydrophobic interaction with edaravone. Interaction of drugs with protein and its mutants may make it possible to restore the stability of SOD1 structure and attenuate disease progression in ALS patients carrying these mutations.
Keywords: Amyotrophic lateral sclerosis (ALS), Cu-Zn superoxide dismutase1 (SOD1), Edaravone, Riluzole, Molecular docking
© 2022 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/)
DOI: http://dx.doi.org/10.22159/ijcr.2022v6i4.207. Journal homepage: https://ijcr.info/index.php/journal
INTRODUCTION
Amyotrophic lateral sclerosis (ALS) is a fatal motor neuron disorder that causes progressive loss of the upper and lower motor neurons in the brain and spinal cord. Individuals who are affected with ALS develop progressive muscle weakness, atrophy and paralysis, which eventually leads to death due to respiratory failure, typically within 2 to 3 y of symptom onset. ALS is categorized into sporadic ALS (sALS), which occurs randomly in the community and is not genetically inherited and familial ALS (fALS), which is genetically inherited through mutated genes. With 90-95% of ALS being sporadic, it is the most common form and the remaining 5–10% of the cases are fALS [1-3]. Both genetics and environment play a role in development of ALS. fALS is caused by mutations in genes like TARDBP, FUS, OPTN, VCP, UBQLN2, C9ORF72, TUBA4A, TDP-43, PFN1 and SOD1 [3-11]. Some of the environmental factors that are associated with ALS are smoking, exposure to chemicals like pesticides, fertilizers, herbicides, insecticides, formaldehyde and metals, exposure to radiation and low-frequency electromagnetic fields that generate a bigger quantity of cellular reactive oxygen than normal, physical activity and diet [2].
Copper-Zinc Superoxide Dismutase 1 (Cu-Zn SOD1) was the first gene to be discovered as the candidate gene for fALS, after linking the disease to a genetic defect on chromosome 21q [12]. SOD1 protein is a ubiquitous soluble protein which is widely distributed in the cytoplasm, nucleus, mitochondrial intermembrane space, peroxisomes and lysosomes [13-16]. A functional SOD1 binds to metals like copper and zinc ions and has an intramolecular disulfide bond connecting C57 and C146, forms a homodimer which catalyses the dismutation reaction of superoxide radicals to hydrogen peroxide and oxygen through the cyclic reduction and reoxidation of copper; acts as a nuclear transcription factor to regulate oxidative stress resistance; plays a role in zinc metabolism; nitration of proteins and repressing respiration being some of the functions among many others [17-22].
A mature dimeric SOD1 protein is extremely stable and is an unlikely candidate for ALS. As ALS is caused by toxic protein misfolding, unfolding and aggregation, it’s the immature monomeric apo forms of SOD1-WT which are unstable and cause cytotoxic misfolded conformations, which are found to be associated with sALS. Mutations in SOD1 dimer are also associated with ALS where more than 150 mutations throughout the sequence of the protein are associated with 5% of sALS and 20 to 25% of fALS. fALS associated SOD1 mutations were thought to lead to a loss of dismutase function, but a toxic gain of function promoted by the formation of misfolded conformers that are prone to aberrant interactions both with each other and with other cellular components also contributes to the disease [22, 23, 29, 44].
Amino acids H64, H72, H81, D84 are found to be involved in zinc ion binding and amino acids H47, H49, H64, H121 are found to be involved in copper ion binding [35, 36]. As metal binding is important for maturation, dimerization and proper functioning of the protein, mutations of some of the amino acids involved in metal binding is studied in this paper. As there are many genetic and protein mutations that can lead to ALS and the only two food and drug administration (FDA) approved drugs for ALS-Edaravone and Riluzole, we used docking studies to determine the interaction of these drugs with the SOD1-WT and SOD1 mutants, to assess if the ALS patients with misfolded SOD1-WT and SOD1 mutants can derive any clinical therapy from the available FDA approved treatment.
MATERIALS AND METHODS
Sequence and structure retrieval of human SOD1
Human SOD1 amino acid sequence was retrieved from UniProt knowledgebase [24] with the search term as human SOD1. The very first result with the UniProt ID P00441 was selected and retrieved in FASTA format. The protein data bank (PDB) structure of human SOD1 was searched in RCSB. org [25] with human SOD1 as search term and a solution structure of the wildtype protein with PDBID 2NAM was retrieved.
Multiple sequence alignment and construction of phylogenetic tree
The FASTA format amino acid sequence of human SOD1 was obtained from NCBI and protein blast was run. 25 protein sequences of other organisms aligned with SOD1 were downloaded in FASTA aligned sequence. Hypothetical proteins, synthetic constructs, ligand-bound proteins, mutated, unnamed, predicted and low-quality proteins were excluded. Phylogenetic tree was constructed using MEGAx, which is an integrated tool for conducting automatic and manual sequence alignment and inferring phylogenetic trees [29]. A new alignment was created in MEGAx and FASTA sequences were inserted and selected, after which clustal was used for the alignment of sequences and the neighbor-joining method was used for the construction of the phylogenetic tree. The amino acids of the metal binding region of the protein whose mutations lead to ALS, were determined for its conservation among all the sequences, as the mutations of the conserved amino acids can significantly distort the tertiary structure of the protein [44].
Prediction of nsSNVs’ effect on SOD1 function
The functional and stability effects of the non-synonymous single nucleotide variants (nsSNVs) of PFN1 were predicted using PredictSNV, which is a dataset consisting of the following algorithms: MAPP, PhD-SNPs, Polyphen1, Polyphen2, SIFT, SNAP, nsSNP Analyzer and PANTHER. PFN1 amino acid sequence from UniProt in FASTA format was loaded in the dataset and the values for each nsSNV was given as confidence percentage for each tool [33]. Four amino acids shown to have mutations causing ALS (H81A, H47R, H49R, H49Q) in the metal binding region of the protein were determined for their effect on the protein function using PredictSNP in which all the tools except nsSNP Analyzer have shown results.
Determination of stereochemical properties of SOD1
Zlab Ramachandran plot was employed for determining the stereochemical properties of SOD1. All the residues were considered except glycine, proline residues and first and last residues of the protein for determination of highly preferred, preferred and questionable conformations in the plot, along with their Phi and Psi angles.
Induction of mutagenesis in SOD1
Out of 5 conformers present in the PDBID 2NAM SOD1 structure, the first one was saved by deleting the rest of the conformers in biovia discovery studio. The saved conformer was loaded in pymol and the positions of the residues to be mutated were selected and changed by applying the mutagenesis wizard and the structures were saved as a PDB file.
ADME analysis of the ligands used for docking studies
ADME analysis is done to determine how a drug is processed in the body. The drugs used for the docking studies are Edaravone and Riluzole, whose drug-likeness was determined using Swiss ADME web tool [34, 35] by inserting the canonical SMILES of the drug, which were retrieved from PubChem [36]. The drug-likeness of the drugs were determined by their Lipinski rule’s parameters and gastrointestinal absorption of the drugs was determined by BOILED-Egg.
Docking studies using autodock suites
To study the interaction of ligands Edaravone and Riluzole with SOD1-WT, SOD1-H81A, SOD1-H49R, autodock suites was used [38]. Autodock is a suite of softwares for predicting the optimal bound conformations of ligands to proteins [42]. Autodock version 4.2.6 was used to predict the conformation and the binding affinity of the ligands Edaravone and Riluzole towards SOD1-WT, SOD1-H81A and SOD1-H49R. The PDB file of the first conformation of SOD1 molecule 2NAM was loaded in the autodock, polar hydrogens and kollman charges were assigned to the molecule and a PDBQT file was created. The 3D conformer SDF file of the ligand was converted to PDB file in PyMOL, which was loaded in autodock and after setting up the rotatable bonds, a PDBQT file of the ligand was created. Grid box was set for blind docking so as to screen the entire protein molecule for active sites and saved the grid parameter file (GPF), which is used for autogrid calculation. Autogrid was launched by using autogrid4 program of the autodock suite, which generates an electrostatic potential grid map and a desolvation map. After defining docking parameters, a docking parameter file (DPF) was generated for autodock calculation. Autodock was launched by using autodock4 program of the autodock suite, which uses the full set of grid maps and the flexible part of the receptor to guide the docking process of the ligands. Docking was done by using genetic algorithm parameters and the free binding energies were obtained based on linear regression analysis in a docking log file (DLG). The best conformation was analysed by the highest binding energies and the respective complex was saved as a PDBQT file which was converted to PDB file in open babel software for further analysis. Protein ligand interaction profiler (PLIP) was used to generate a three-dimensional fig. of the protein-ligand complex interaction determining the site of interaction between the protein and ligand [43].
RESULTS
Sequence and structure retrieval of human SOD1
Uniprot Knowledgebase search was performed with the search term as human SOD1, which displayed 135 results out of which only first four results were of SOD1 from different organisms. The very first result, which displayed SOD1 amino acid sequence of Homo sapiens with the UniProt ID P00441 was selected and retrieved in FASTA format. SOD1 is a 16KDa actin-binding protein consisting of 154 amino acids [24] coded by SOD1 gene which is located on chromosome 21q22.11 [26]. Protein database (PDB) searches were performed in RCSB. Org with human SOD1 as search term and the monomeric structure with PDBID 2NAM was retrieved which is a solution structure of a full-length SOD1-WT.
PDBID-2NAM SOD1 structure
The three-dimensional solution structure of the SOD1 molecule was determined by solution nuclear magnetic resonance (NMR) spectroscopy [27]. This molecule has an ensemble of five conformational structures, out of which we used the very first structure by deleting the rest of the structural conformations using Biovia Discovery studio and saving the first conformation in PDB format which was used for this study. The saved first conformer was loaded in PyMOL and saved in Portable network graphics (PNG) format for image (fig. 1).
In a fully functional mature homodimeric SOD1 protein, each subunit folds into an eight-stranded, Greek-key β-barrel and holds one copper and one zinc ion, where the homodimer is further stabilized by an intramolecular disulfide bridge Cys57-Cys146 [28]. But a nascent WT SOD1 (PDBID-2NAM) without binding to ions and without a disulfide bridge is highly disordered, without any β-sheet structure, as it requires binding to ions to form a well-folded β-structure [29].
Fig. 1: PDBID-2NAM, the monomeric solution structure of the first conformer of full-length human wildtype Copper-Zinc Superoxide dismutase1 (SOD1). The image was saved from PyMOL
Multiple sequence alignment and construction of phylogenetic tree using MEGAx
Multiple sequence alignment and phylogenetic tree was constructed using MEGAx, which is an integrated tool that can be used for conducting automatic and manual sequence alignment and inferring phylogenetic trees [33]. FASTA format sequence of SOD1 retrieved from Uniprot was loaded and a protein blast was run. 25 protein sequences of other organisms aligned with SOD1 were downloaded in FASTA aligned sequence. In MEGAx, a new alignment was created and after the FASTA sequences were inserted and selected, they were aligned by using ClustalW. The phylogenetic tree was made using the neighbor-joining method (fig. 2). Among the four mutations H81A, H47R, H49R, H49Q in the metal binding region, H81A, H47R and H49Q were found to be in conserved regions, which can be more deleterious and can significantly distort the tertiary structure of the protein [44].
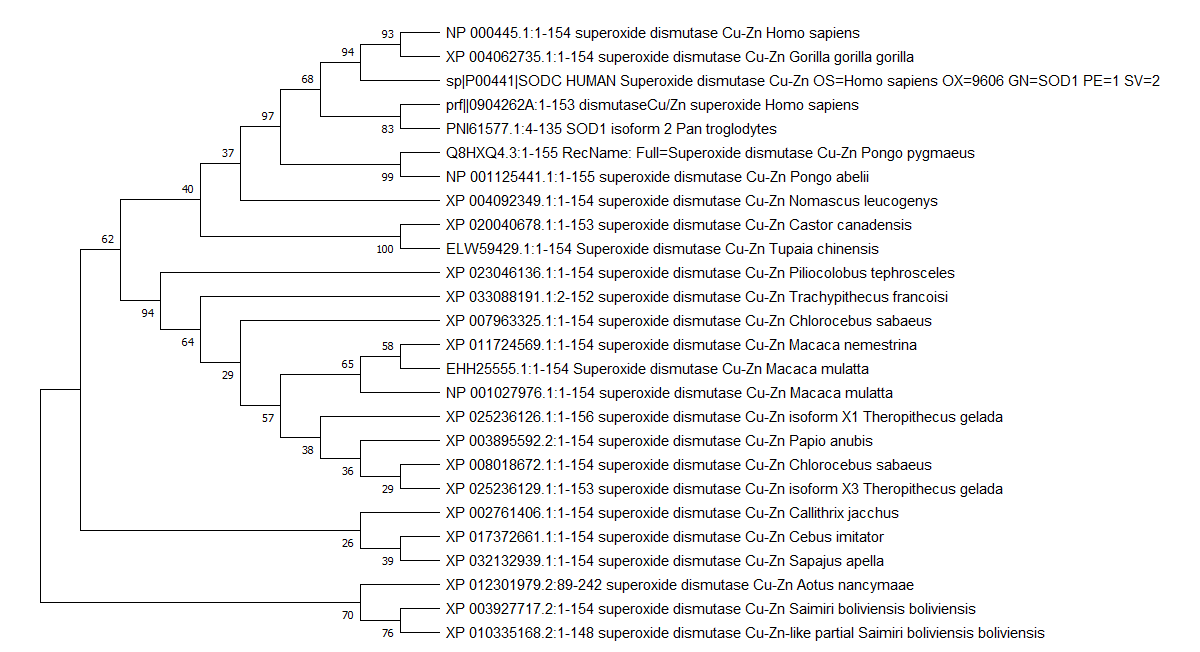
Fig. 2: The evolutionary history was inferred using the Neighbor-Joining method [30]. The bootstrap consensus tree inferred from 500 replicates is taken to represent the evolutionary history of the taxa analyzed. Branches corresponding to partitions reproduced in less than 50% bootstrap replicates are collapsed. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (500 replicates) are shown next to the branches [31]. The evolutionary distances were computed using the Poisson correction method [32] and are in the units of the number of amino acid substitutions per site. This analysis involved 26 amino acid sequences. All ambiguous positions were removed for each sequence pair (pairwise deletion option). There was a total of 158 positions in the final dataset. Evolutionary analyses were conducted in MEGA X [33]
Prediction of nsSNVs’ effect on protein function using PredictSNP
The single nucleotide variants (SNVs) cause the most frequent type of genetic variations in humans, out of which some changes are neutral and do not cause any adverse effect on protein function, while others affect the gene expression or the function of the translated protein which leads to the development of various diseases. Approximately half of the known disease-related mutations stems from non-synonymous SNVs (nsSNVs), manifested as amino acid mutations. It is important to know the association between SNVs and its related diseases, but it is difficult to distinguish pathogenic substitutions from those that are functionally neutral by any experimental assay. Computational prediction tools like PredictSNP are valuable for the initial analysis of SNVs and their effect on protein function. PredictSNP is a dataset containing eight established prediction tools MAPP, nsSNPAnalyzer, PANTHER, PhD-SNP, PolyPhen-1, PolyPhen-2, SIFT and SNAP [34]. Out of eight amino acids that are involved in metal binding (H64, H72, H81, D84 for zinc ion binding, H47, H49, H64, H121 for copper ion binding), four amino acids shown to have mutations causing ALS (H81A, H47R, H49R, H49Q) from Uniprot [24] were determined for their effect on the protein function using PredictSNP in which all the tools except nsSNPAnalyzer have shown results. All the mutations were found to be deleterious with a confidence percentage of 84% (table 1). H81A mutation in the conserved region, which inhibits zinc binding and H49R mutation in a non-conserved region, which inhibits copper binding, were considered for this study.
Table 1: Functional effect of amino acid mutations involved in metal binding in SOD1
| Mutation | PredictSNP | MAPP | PHD-SNP | Polyphen1 | Polyphen2 | SIFT | SNAP | Panther |
| H47R | 87% | 77% | 86% | 74% | 68% | 53% | 89% | 72% |
| H49Q | 87% | 77% | 82% | 74% | 68% | 79% | 89% | 77% |
| H49R | 87% | 92% | 86% | 74% | 59% | 79% | 89% | 65% |
| H81A | 87% | 88% | 82% | 74% | 81% | 79% | 85% | 72% |
Determination of stereochemical properties of the protein using Zlab
Zhang lab Ramachandran plot server was employed for obtaining the stereochemical properties of SOD1 [37]. The plot was obtained by considering a total of 122 residues leaving the glycine and proline residues and the first residue of N-termini and the last residue of C-termini. 99 residues which makes 81.148% were shown to have highly preferred conformations. 18 residues which makes 14.754% were shown to have preferred conformations and 5 residues which makes 4.098% were shown to have questionable conformations. H81A had a Phi angle of-80.95 and a psi angle of-41.74 and was shown to have a highly preferred conformation where delta>=-2. H49R had a Phi angle of-91.68 and a Psi angle of-47.3 was shown to have a highly preferred conformation where delta>=-2 (fig. 3).
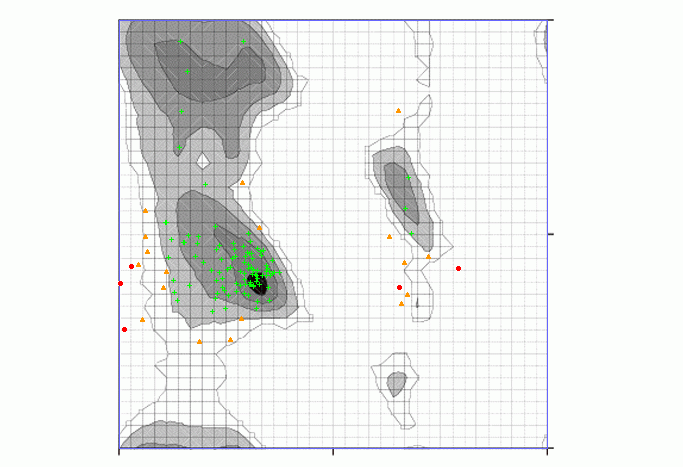
Fig. 3: Ramachandran plot obtained using Zlab. Black, dark grey, grey, light grey represent highly preferred conformations where delta>=-2, white with black grid represents preferred conformations where-2>delta>-4 and white with grey grid represents questionable conformations where delta<-4. 81.148% of residues have highly preferred conformations shown as green crosses, 14.754% of residues have preferred conformations shown as brown triangles and 4.098% of the residues have questionable conformations shown as red circles
Inducing mutagenesis in SOD1 using PyMOL
Mutagenesis was induced in PyMOL to create nsSNPs by loading the first conformational structure saved from biovia discovery studio, selecting the residue at the position to be mutated and applying the mutagenesis wizard. The histidine residue which is a polar amino acid having an aromatic ring was changed to alanine which is a small, aliphatic and non-polar amino acid having a hydrophobic side chain. The histidine residue at the 49th position is substituted with arginine, which is a positively charged, hydrophilic amino acid (fig. 4).
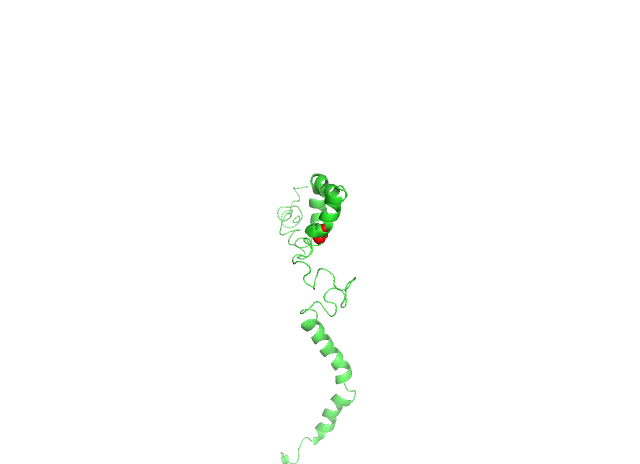
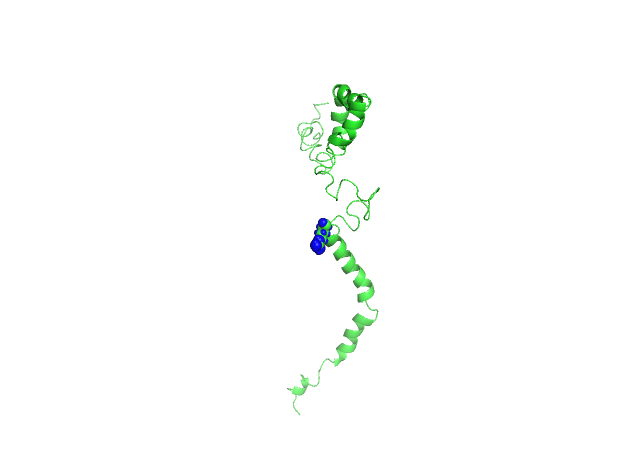
Fig. 4: 4A-Histidine residue at 81st position is changed to alanine (H81A) shown as red sphere, 4B-Histidine residue at 49th position is changed to arginine (H49R) shown as a blue sphere. The structure is represented in the ribbon form and the SNV is represented in the sphere form
ADME analysis of the ligands used for docking studies
The two FDA-approved drugs for ALS, Edaravone and Riluzole were considered to be tested for their interaction with both wild-type and mutated SOD1. The drug-likeness of the ligands were determined using SwissADME web tool by inserting the canonical SMILES obtained from PubChem [38-40]. Edaravone is a free radical scavenger used to delay the progression of ALS. It has a molecular formula C10H10N2O with a molecular weight of 174.20g/mol. It follows the lipinski rule as it has high gastrointestinal absorption and is blood brain barrier permeant as shown by boiled egg; the number of hydrogen bond acceptors and donors for edaravone being 2 and 0 respectively and the lipophilicity being 2.07. Riluzole is a glutamate antagonist, used as anticonvulsant in the treatment of ALS, to prolong the survival of patients [41]. It has a molecular formula of C8H5F3N2O5 with a molecular weight of 234.20g/mol. It follows lipinski rule as it has high gastrointestinal absorption and is blood-brain barrier permeant, as shown by boiled egg; the number of hydrogen bond acceptors and donors for riluzole being 5 and 1, respectively and the lipophilicity being 2.04. The three-dimensional structures of both the drugs were retrieved from Pubchem (fig. 5).
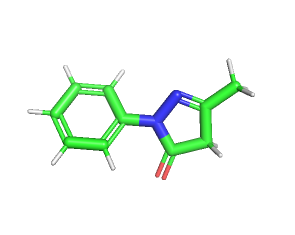
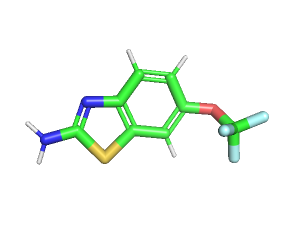
Fig. 5: The three-dimensional ball and stick model structures were retrieved from Pubchem. 5A-Edaravone, 5B-Riluzole
Docking studies of wildtype and mutant (H81A, H49R) SOD1 with edaravone and riluzole in autodock
Autodock is a suite of softwares for predicting the optimal bound conformations of ligands to proteins [42]. Autodock version 4.2.6 was used to predict the conformation and the binding affinity of the ligands edaravone and riluzole towards the SOD1-WT, SOD1-H81A and SOD1-H49R. The PDB file of the first conformation of SOD1 molecule 2NAM was loaded in the autodock, polar hydrogens and kollman charges were assigned to the molecule and a PDBQT file was created. The 3D conformer SDF file of the ligand was converted to PDB file in PyMOL which was loaded in autodock and after setting up the rotatable bonds, a PDBQT file of the ligand was created. Grid box was set for blind docking so as to screen the entire protein molecule for active sites and saved the grid parameter file (GPF), which is used for autogrid calculation. Autogrid was launched by using autogrid4 program of the autodock suite, which generates an electrostatic potential grid map and a desolvation map. After defining docking parameters, a docking parameter file (DPF) was generated for autodock calculation. Autodock was launched by using autodock4 program of the autodock suite, which uses the full set of grid maps and the flexible part of the receptor to guide the docking process of the ligands. Docking was done by using genetic algorithm parameters and the free binding energies were obtained based on linear regression analysis in a docking log file (DLG).
The ligands were found to interact with the wild type as well as SOD1 mutants. The best conformation was analysed by the highest binding energies (table 2) and the complex was saved as a PDBQT file which was converted to PDB file in open babel software for further analysis. Protein-ligand interaction profiler (PLIP) was used to generate a three-dimensional fig. of the protein-ligand complex interaction determining the site of interaction between the protein and ligand [43].
SOD1-WT and mutants interact at different sites with both drugs. SOD1-WT was found to interact at V103, H110, R115 through hydrophobic interactions and at H110 through hydrogen bond with edaravone. SOD1-WT was found to interact at T54, F64 through hydrophobic interactions and at D52, T58 through hydrogen bond with riluzole (fig. 6). SOD1-H81A was found to interact at L38, L42, F45 through hydrophobic interactions and at H46 through hydrogen bond with edaravone. SOD1-H81A was found to interact at F45, D52 through hydrophobic interactions and at G37, H46 through hydrogen bond with riluzole (fig. 7). SOD1-H49R was found to interact at L38, L42 with hydrophobic interactions and at H46 through a hydrogen bond with edaravone. SOD1-H49R was found to interact at F45 through hydrophobic interactions and at H46 through a hydrogen bond with riluzole (fig. 8).
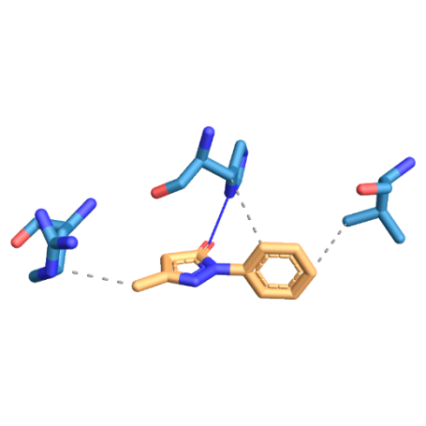
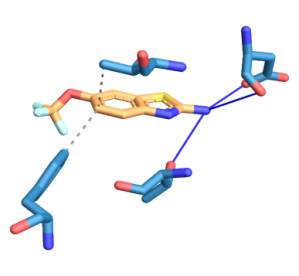
Fig. 6: Three-dimensional figure of the protein-ligand complex were generated by protein ligand interaction profiler. The hydrophobic (dotted lines) and hydrogen bond (solid blue line) interactions between the ligand and the protein SOD1. 6A: SOD1-WT was found to interact at V103, H110, R115 through hydrophobic interactions and at H110 through hydrogen bond with edaravone. 6B: SOD1-WT was found to interact at T54, F64 through hydrophobic interactions and at D52, T58 through hydrogen bond with riluzole
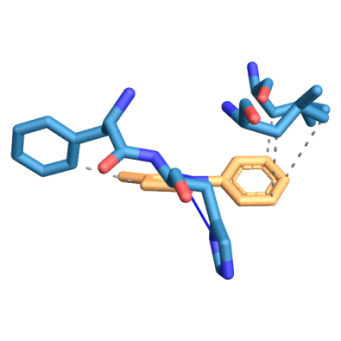
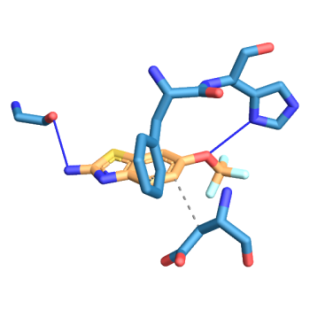
Fig. 7: Three-dimensional figure of the protein-ligand complex were generated by protein ligand interaction profiler. The hydrophobic (dotted lines) and hydrogen bond (solid blue line) interactions between the ligand and the protein SOD1-H81A. 7A: SOD1-H81A was found to interact at L38, L42, F45 through hydrophobic interactions and at H46 through a hydrogen bond with edaravone. 7B: SOD1-H81A was found to interact at F45, D52 through hydrophobic interactions and at G37, H46 through hydrogen bond with riluzole
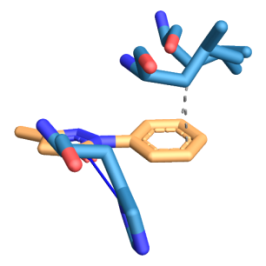
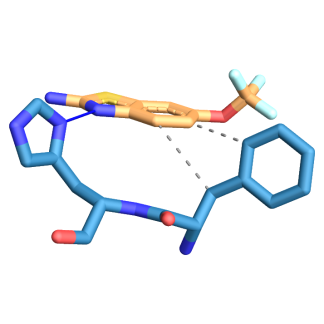
Fig. 8: Three-dimensional figure of the protein-ligand complex were generated by protein ligand interaction profiler. The hydrophobic (dotted lines) and hydrogen bond (solid blue line) interactions between the ligand and the protein SOD1-H81A. 8A: SOD1-H49R was found to interact at L38, L42 with hydrophobic interactions and at H46 through hydrogen bond with edaravone. 8B: SOD1-H49R was found to interact at F45 through hydrophobic interactions and at H46 through hydrogen bond with riluzole
Table 2: The binding energies obtained from docking of wild-type and mutant SOD1 with edaravone and riluzole
| S. No. | Protein | Ligand | Reference RMSD | Binding energy (Kcal/mol) | Inhibition constant (mM) |
| 1. | SOD1-WT | Edaravone | 206.30 | -3.25 | 4.16 |
| 2. | SOD1-WT | Riluzole | 176.79 | -2.95 | 6.89 |
| 3 | SOD1-H81A | Edaravone | 170.88 | -3.45 | 2.96 |
| 4. | SOD1-H81A | Riluzole | 170.15 | -2.76 | 9.54 |
| 5. | SOD1-H49R | Edaravone | 170.68 | -3.49 | 2.75 |
| 6. | SOD1-H49R | Riluzole | 171.11 | -2.88 | 7.74 |
DISCUSSION
ALS is a progressive condition caused by the deterioration of the motor neurons present in the spinal cord and brain, resulting in paralysis. The survival of patients with ALS is 2 to 5 y; regardless of this glum prognosis, 20% of patients live for 5 y, 10% for 10 y, and 5% for 20 y or longer. In 1993, SOD1 was the first gene linked to ALS, in which more than 150 mutations throughout the sequence of the protein are associated with 5% of sALS and 20 to 25% of fALS cases [12, 22, 47].
A mature dimeric SOD1 protein requires binding to a Cu and Zn ions and an intramolecular disulfide bond connecting C57 and C146 residues. Amino acids H64, H72, H81, D84 are involved in zinc ion binding and H47, H49, H64, H121 are involved in copper ion binding [35, 36]. The zinc ion stabilizes the protein, promotes homodimerization and softens the active site so that copper cycling between Cu(I) and Cu(II) can rapidly occur, whereas the copper ion is crucial to the enzymatic activity of SOD1 [46, 52]. And so, SOD1 metal binding mutants remain immature, which over time induces pathological effects in the cell by exhibiting pro-oxidant activity [52, 53, 55]. The nascent SOD1-WT becomes highly disordered in the absence of metal ions, especially zinc metal ion [29].
Both monomeric apo SOD1-WT and SOD1 dimer metal binding region (MBR) mutations are found to cause ALS. In this paper, we studied the MBR mutations of monomeric apo SOD1-WT protein molecule as even the mutations in MBR of SOD1 dimer impairs the dimerization of the protein molecule and promotes monomer formation. This apo SOD1-WT and SOD1 mutant monomers undergo unfolding, misfolding and gain of cytotoxicity by acquiring a novel capacity of interacting with membranes and form aggregates. Aggregation also occurs by forming heterodimers and oligomers between both type of monomers which decreases antioxidant activity, increases oxidative damage, reduces longevity and oxidative stress-induced mutant SOD1 aggregation, which might represent the toxic precursor species in the cell for ALS [48, 50, 54].
In a functional SOD1 dimer, the β-barrel is normally protected against self-aggregation by having each end of the dimer covered by electrostatic and Zn-binding loops, which becomes disordered in the SOD1 metal binding site mutants through loss of metal, exposing the β-barrels of the dimer to the external environment, allowing close interactions with adjacent molecules, that causes misfolding and aggregation of the protein. These observations support the hypothesis that the unfolded form of SOD1, after synthesis on the ribosome and prior to folding and maturation, is the source of misfolding and aggregation in motor neurons associated with sALS [45, 46, 55].
ALS patients carrying mutations in other genes such as C9orf72HRE, FUS, KIF5A, NEK1, VAPB and ALSIN are also found with misfolded SOD1-WT positive intracytoplasmic inclusions in motor neurons, suggesting that misfolding of SOD1-WT can be part of a common downstream event that may be pathogenic [51]. The unfavorable conformational changes [57] in SOD1 protein may not interact with other associated molecules, which may block the mediated functions of downstream molecules [44].
Four amino acid mutations involved in metal binding were retrieved from Uniprot (H81A, H47R, H49R and H49Q) and two mutations which were in conserved regions; H81A involved in Zn binding and H49R involved in Cu binding, were considered for this study as mutations in a conserved domain of SOD1 protein are more deleterious and significantly distort the tertiary structure of the protein (44). For the docking studies, PDBID 2NAM, the monomeric human SOD1-WT solution structure, was considered, as the process of crystallography can exert pressure on the proteins due to reduction in humidity and temperature change which can effect the actual structure of the protein and the solution structure’s microenvironment being closer to the natural microenvironment [59].
It took 22 y for the FDA to approve Edaravone in 2017 after approving Riluzole in 1995 [60]. Even though their mechanism of action is not known, they have shown a significant positive effect on the rates of survival, muscle deterioration [61] and the long-term efficacy in the clinical trials for the treatment of ALS [62]. These two drugs were docked with SOD1-WT and SOD1 mutants (H81A, H49R) to determine the type of interaction, degree of interaction and their binding energies. SOD1-WT has hydrophobic interactions at V103, H110, R115 and a hydrogen bond at H110 with Edaravone. SOD1-WT has hydrophobic interactions at T54, F64 and hydrogen bond at D52, T58 with Riluzole. Both SOD1 mutants has a hydrogen bond at its H46 residue for both the drugs. SOD1-H49R mutant has hydrophobic interactions at F45 with Riluzole. SOD1-H81A mutant has hydrophobic interactions at F45 with Edaravone. For both the mutants L42A residue has a hydrophobic interaction with Edaravone.
CONCLUSION
As the drugs bind to ALS, causing mutated SOD1 protein, it can be possible to restore the stability of SOD1 structure due to the interaction of these drugs and disease progression can be attenuated in the ALS patients carrying these mutations. Other ALS-causing mutations in SOD1 and other proteins can be studied for their interaction with drugs that can be informative in preclinical and clinical studies for the development of a novel therapy for ALS patients.
ACKNOWLEDGEMENT
We gratefully acknowledge Prof. S S Singh, Dept. of Biochemistry, Osmania University, for his support while writing the manuscript. I extend my gratitude to BioNome, India, their research team and staff for their valuable guidance and support in the completion of the project.
FUNDING
Nil
AUTHORS CONTRIBUTIONS
All the authors have contributed equally.
CONFLICT OF INTERESTS
Declared none
REFERENCES
-
Fil D, DeLoach A, Yadav S, Alkam D, MacNicol M, Singh A. Mutant profilin1 transgenic mice recapitulate cardinal features of motor neuron disease. Hum Mol Genet. 2017 Feb 15;26(4):686-701. doi: 10.1093/hmg/ddw429, PMID 28040732.
-
Zarei S, Carr K, Reiley L, Diaz K, Guerra O, Altamirano PF. A comprehensive review of amyotrophic lateral sclerosis. Surg Neurol Int. 2015;6:171. doi: 10.4103/2152-7806.169561, PMID 26629397.
-
Renton AE, Chio A, Traynor BJ. State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci. 2014 Jan;17(1):17-23. doi: 10.1038/nn.3584, PMID 24369373.
-
Van Blitterswijk M, DeJesus Hernandez M, Rademakers R. How do C9ORF72 repeat expansions cause amyotrophic lateral sclerosis and frontotemporal dementia: can we learn from other noncoding repeat expansion disorders? Curr Opin Neurol. 2012 Dec 1;25(6):689-700. doi: 10.1097/WCO.0b013e32835a3efb, PMID 23160421.
-
Abramzon Y, Johnson JO, Scholz SW, Taylor JP, Brunetti M, Calvo A. Valosin-containing protein (VCP) mutations in sporadic amyotrophic lateral sclerosis. Neurobiol Aging. 2012;33(9):2231.e1-6. doi: 10.1016/j.neurobiolaging.2012.04.005. PMID 22572540.
-
Smith BN, Ticozzi N, Fallini C, Gkazi AS, Topp S, Kenna KP. Exome-wide rare variant analysis identifies TUBA4A mutations associated with familial ALS. Neuron. 2014 Oct 22;84(2):324-31. doi: 10.1016/j.neuron.2014.09.027, PMID 25374358.
-
Bruijn LI, Becher MW, Lee MK, Anderson KL, Jenkins NA, Copeland NG. ALS-linked SOD1 mutant G85R mediates damage to astrocytes and promotes rapidly progressive disease with SOD1-containing inclusions. Neuron. 1997 Feb 1;18(2):327-38. doi: 10.1016/s0896-6273(00)80272-x, PMID 9052802.
-
Ripps ME, Huntley GW, Hof PR, Morrison JH, Gordon JW. Transgenic mice expressing an altered murine superoxide dismutase gene provide an animal model of amyotrophic lateral sclerosis. Proc Natl Acad Sci USA. 1995 Jan 31;92(3):689-93. doi: 10.1073/pnas.92.3.689, PMID 7846037.
-
Wegorzewska I, Bell S, Cairns NJ, Miller TM, Baloh RH. TDP-43 mutant transgenic mice develop features of ALS and frontotemporal lobar degeneration. Proc Natl Acad Sci USA. 2009 Nov 3;106(44):18809-14. doi: 10.1073/pnas.0908767106, PMID 19833869.
-
Ingre C, Landers JE, Rizik N, Volk AE, Akimoto C, Birve A. A novel phosphorylation site mutation in profilin 1 was revealed in a large screen of US, Nordic, and German amyotrophic lateral sclerosis/frontotemporal dementia cohorts. Neurobiol Aging. 2013;34(6):1708.e1-6. doi: 10.1016/j.neurobiolaging.2012.10.009. PMID 23141414.
-
Smith BN, Vance C, Scotter EL, Troakes C, Wong CH, Topp S. Novel mutations support a role for profilin 1 in the pathogenesis of ALS. Neurobiol Aging. 2015;36(3):1602.e17-27. doi: 10.1016/j.neurobiolaging.2014.10.032. PMID 25499087.
-
Rosen DR, Siddique T, Patterson D, Figlewicz DA, Sapp P, Hentati A. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature. 1993 Mar;362(6415):59-62. doi: 10.1038/362059a0, PMID 8446170.
-
Chang LY, Slot JW, Geuze HJ, Crapo JD. Molecular immunocytochemistry of the CuZn superoxide dismutase in rat hepatocytes. J Cell Biol. 1988;107(6 Pt 1):2169-79. doi: 10.1083/jcb.107.6.2169, PMID 3058718.
-
Keller GA, Warner TG, Steimer KS, Hallewell RA. Cu, Zn superoxide dismutase is a peroxisomal enzyme in human fibroblasts and hepatoma cells. Proc Natl Acad Sci USA. 1991 Aug 15;88(16):7381-5. doi: 10.1073/pnas.88.16.7381, PMID 1651504.
-
Crapo JD, Oury T, Rabouille C, Slot JW, Chang LY. Copper, zinc superoxide dismutase is primarily a cytosolic protein in human cells. Proc Natl Acad Sci USA. 1992 Nov 1;89(21):10405-9. doi: 10.1073/pnas.89.21.10405, PMID 1332049.
-
Sturtz LA, Diekert K, Jensen LT, Lill R, Culotta VC. A fraction of yeast Cu, Zn-superoxide dismutase and its metallochaperone, CCS, localize to the intermembrane space of mitochondria. A physiological role for SOD1 in guarding against mitochondrial oxidative damage. J Biol Chem. 2001 Oct 12;276(41):38084-9. doi: 10.1074/jbc.M105296200. PMID 11500508.
-
Reddi AR, Culotta VC. SOD1 integrates signals from oxygen and glucose to repress respiration. Cell. 2013 Jan 17;152(1-2):224-35. doi: 10.1016/j.cell.2012.11.046, PMID 23332757.
-
Tsang CK, Liu Y, Thomas J, Zhang Y, Zheng XF. Superoxide dismutase 1 acts as a nuclear transcription factor to regulate oxidative stress resistance. Nat Commun. 2014 Mar 19;5(1):3446. doi: 10.1038/ncomms4446, PMID 24647101.
-
Wei JP, Srinivasan C, Han H, Valentine JS, Gralla EB. Evidence for a novel role of copper-zinc superoxide dismutase in zinc metabolism. J Biol Chem. 2001 Nov 30;276(48):44798-803. doi: 10.1074/jbc.M104708200. PMID 11581253.
-
Beckman JS, Carson M, Smith CD, Koppenol WH. ALS, SOD and peroxynitrite. Nature. 1993 Aug;364(6438):584. doi: 10.1038/364584a0, PMID 8350919.
-
Tainer JA, Getzoff ED, Richardson JS, Richardson DC. Structure and mechanism of copper, zinc superoxide dismutase. Nature. 1983 Nov;306(5940):284-7. doi: 10.1038/306284a0, PMID 6316150.
-
Saccon RA, Bunton Stasyshyn RK, Fisher EM, Fratta P. Is SOD1 loss of function involved in amyotrophic lateral sclerosis? Brain. 2013 Aug 1;136(8):2342-58. doi: 10.1093/brain/awt097, PMID 23687121.
-
Culik RM, Sekhar A, Nagesh J, Deol H, Rumfeldt JAO, Meiering EM. Effects of maturation on the conformational free-energy landscape of SOD1. Proc Natl Acad Sci USA. 2018 Mar 13;115(11):E2546-55. doi: 10.1073/pnas.1721022115, PMID 29483249.
-
The UniProt Consortium. UniProt: the universal protein KnowledgeBase. Nucleic Acids Res. 2017;45(D1):D158-69. doi: 10.1093/nar/gkw1099, PMID 27899622.
-
Rose PW, Prlić A, Altunkaya A, Bi C, Bradley AR, Christie CH. The RCSB protein data bank: an integrative view of protein, gene and 3D structural information. Nucleic Acids Res. 2017;45(D1):D271-81. doi: 10.1093/nar/gkw1000, PMID 27794042.
-
Amberger JS, Bocchini CA, Scott AF, Hamosh A. OMIM.org: leveraging knowledge across phenotype-gene relationships. Nucleic Acids Res. 2019;47(D1):D1038-43. doi: 10.1093/nar/gky1151. PMID 30445645.
-
Puthenveetil R, Vinogradova O. Solution NMR: A powerful tool for structural and functional studies of membrane proteins in reconstituted environments. J Biol Chem. 2019 Nov 1;294(44):15914-31. doi: 10.1074/jbc.REV119.009178. PMID 31551353.
-
Parge HE, Hallewell RA, Tainer JA. Atomic structures of wild-type and thermostable mutant recombinant human Cu, Zn superoxide dismutase. Proc Natl Acad Sci USA. 1992 Jul 1;89(13):6109-13. doi: 10.1073/pnas.89.13.6109, PMID 1463506.
-
Lim L, Song J. SALS-linked WT-SOD1 adopts a highly similar helical conformation as FALS-causing L126Z-SOD1 in a membrane environment. Biochim Biophys Acta. 2016;1858(9):2223-30. doi: 10.1016/j.bbamem.2016.06.027. PMID 27378311.
-
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul 1;4(4):406-25. doi: 10.1093/oxford journals.molbev.a040454. PMID 3447015.
-
Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985 Jul;39(4):783-91. doi: 10.1111/j.1558-5646.1985.tb00420.x. PMID 28561359.
-
Zuckerkandl E, Pauling L. Evolutionary divergence and convergence in proteins. Inevolving genes and proteins. Academic Press; 1965 Jan 1. p. 97-166.
-
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018 Jun;35(6):1547-9. doi: 10.1093/molbev/msy096, PMID 29722887.
-
Bendl J, Stourac J, Salanda O, Pavelka A, Wieben ED, Zendulka J. PredictSNP: robust and accurate consensus classifier for prediction of disease-related mutations. PLoS Comput Biol. 2014 Jan 16;10(1):e1003440. doi: 10.1371/journal.pcbi.1003440. PMID 24453961.
-
Seetharaman SV, Taylor AB, Holloway S, Hart PJ. Structures of mouse SOD1 and human/mouse SOD1 chimeras. Arch Biochem Biophys. 2010 Nov 15;503(2):183-90. doi: 10.1016/j.abb.2010.08.014, PMID 20727846.
-
Nordlund A, Leinartaitė L, Saraboji K, Aisenbrey C, Grobner G, Zetterstrom P. Functional features cause misfolding of the ALS-provoking enzyme SOD1. Proc Natl Acad Sci USA. 2009 Jun 16;106(24):9667-72. doi: 10.1073/pnas.0812046106, PMID 19497878.
-
Anderson RJ, Weng Z, Campbell RK, Jiang X. Main‐chain conformational tendencies of amino acids. Proteins. 2005 Sep 1;60(4):679-89. doi: 10.1002/prot.20530, PMID 16021632.
-
Doogue MP, Polasek TM. The ABCD of clinical pharmacokinetics. Ther Adv Drug Saf. 2013 Feb;4(1):5-7. doi: 10.1177/2042098612469335, PMID 25083246.
-
Daina A, Michielin O, Zoete V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. 2017 Mar 3;7(1):42717. doi: 10.1038/srep42717, PMID 28256516.
-
Kim S, Chen J, Cheng T, Gindulyte A, He J, He S. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Research. 2021 Jan 8;49(D1):D1388-95. https://doi.org/10.1093/nar/gkaa971, PMID 33151290.
-
Jaiswal MK. Riluzole and edaravone: A tale of two amyotrophic lateral sclerosis drugs. Medicinal Research Reviews. 2019 Mar;39(2):733-48. https://doi.org/10.1002/med.21528, PMID 30101496.
-
Forli S, Huey R, Pique ME, Sanner MF, Goodsell DS, Olson AJ. Computational protein–ligand docking and virtual drug screening with the AutoDock suite. Nature Protocols. 2016 May;11(5):905-19. https://doi.org/10.1038/nprot.2016.051, PMID 27077332.
-
Salentin S, Schreiber S, Haupt VJ, Adasme MF, Schroeder M. PLIP: fully automated protein–ligand interaction profiler. Nucleic Acids Research. 2015 Jul 1;43(W1):W443-7. https://doi.org/10.1093/nar/gkv315, PMID 25873628.
-
Pal S, Tiwari A, Sharma K, Sharma SK. Does conserved domain SOD1 mutation has any role in ALS severity and therapeutic outcome? BMC Neuroscience. 2020 Dec;21(1):1-7. https://doi:42.org/10.1186/s12868-020-00591-3, PMID 33036560.
-
Strange RW, Yong CW, Smith W, Hasnain SS. Molecular dynamics using atomic-resolution structure reveal structural fluctuations that may lead to polymerization of human Cu–Zn superoxide dismutase. Proceedings of the National Academy of Sciences USA. 2007 Jun 12;104(24):10040-4. https://doi.org/10.1073/pnas.0703857104.
-
Cohen NR, Zitzewitz JA, Bilsel O, Matthews CR. Nonnative structure in a peptide model of the unfolded state of superoxide dismutase 1 (SOD1): Iimplications for ALS-linked aggregation. Journal of Biological Chemistry. 2019 Sep 13;294(37):13708-17. https://doi.org/10.1074/jbc. PMID 31341015.
-
Hulisz D. Amyotrophic lateral sclerosis: Disease state overview. Am J Manag Care. 2018;24(15)Suppl:S320-6. PMID 30207670.
-
Banci L, Bertini I, Boca M, Girotto S, Martinelli M, Valentine JS, Vieru M. SOD1 and amyotrophic lateral sclerosis: mutations and oligomerization. PloS ONE. 2008 Feb 27;3(2):e1677. https://doi.org/10.1371/journal. PMID 18301754.
-
McCampbell A, Cole T, Wegener AJ, Tomassy GS, Setnicka A, Farley BJ, Schoch KM, Hoye ML, Shabsovich M, Sun L, Luo Y. Antisense oligonucleotides extend survival and reverse decrement in muscle response in ALS models. The Journal of Clinical Investigation. 2018 Nov 1;128(8):3558-67. https://doi.org/10.1172/JCI99081, PMID 30010620.
-
de Araujo Brasil AA, de Carvalho MDC, Gerhardt E, Queiroz DD, Pereira MD, Outeiro TF. Characterization of the activity, aggregation, and toxicity of heterodimers of WT and ALS-associated mutant Sod1. Proceedings of the National Academy of Sciences USA. 2019 Dec 17;116(51):25991-6000. https://doi.org/10.1073/pnas.1902483116, PMID 31796595.
-
Forsberg K, Graffmo K, Pakkenberg B, Weber M, Nielsen M, Marklund S. Misfolded SOD1 inclusions in patients with mutations in C9orf72 and other ALS/FTD-associated genes. Journal of Neurology, Neurosurgery and Psychiatry. 2019 Aug 1;90(8):861-9. https://doi.org/10.1136/jnnp-2018-319386, PMID 30992335.
-
Boyd SD, Ullrich MS, Calvo JS, Behnia F, Meloni G, Winkler DD. Mutations in superoxide dismutase 1 (Sod1) linked to familial amyotrophic lateral sclerosis can disrupt high-affinity zinc-binding promoted by the copper chaperone for Sod1 (Ccs). Molecules. 2020 Jan;25(5):1086. https://doi.org/10.3390/molecules25051086, PMID 32121118.
-
Nagao C, Kuroi K, Wakabayashi T, Nakabayashi T. Pro-oxidant activity of an ALS-linked SOD1 mutant in Zn-deficient form. Molecules. 2020 Jan;25(16):3600. https://doi.org/10.3390/molecules25163600, PMID 32784718.
-
Kim J, Lee H, Lee JH, Kwon DY, Genovesio A, Fenistein D. Dimerization, oligomerization, and aggregation of human amyotrophic lateral sclerosis copper/zinc superoxide dismutase 1 protein mutant forms in live cells. Journal of Biological Chemistry. 2014 May 23;289(21):15094-103. https://doi.org/10.1074/jbc, PMID 24692554.
-
Prudencio M, Borchelt DR. Superoxide dismutase 1 encoding mutations linked to ALS adopts a spectrum of misfolded states. Molecular Neurodegeneration. 2011 Dec;6(1):1-9. https://doi:77.org/10.1186/1750-1326-6-77, PMID 22094223.
-
Borel F, Gernoux G, Cardozo B, Metterville JP, Toro Cabrera GC, Song L. Therapeutic rAAVrh10 mediated SOD1 silencing in adult SOD1(G93A) mice and nonhuman primates. Hum Gene Ther. 2016;27(1):19-31. doi: 10.1089/hum.2015.122. PMID 26710998.
-
Roberts BR, Tainer JA, Getzoff ED, Malencik DA, Anderson SR, Bomben VC, Meyers KR, Karplus PA, Beckman JS. Structural characterization of zinc-deficient human superoxide dismutase and implications for ALS. Journal of Molecular Biology. 2007 Nov 2;373(4):877-90. doi: 10.1016/j.jmb.2007.07.043. PMID 17888947.
-
Wong PC, Pardo CA, Borchelt DR, Lee MK, Copeland NG, Jenkins NA. An adverse property of a familial ALS-linked SOD1 mutation causes motor neuron disease characterized by vacuolar degeneration of mitochondria. Neuron. 1995 Jun 1;14(6):1105-16. https://doi.org/10.1016/0896-6273(95)90259-7.
-
Sadr AS, Eslahchi C, Ghassempour A, Kiaei M. In silico studies reveal structural deviations of mutant profilin-1 and interaction with riluzole and edaravone in amyotrophic lateral sclerosis. Scientific Reports. 2021 Mar 25;11(1):1-4. https://doi:6849.org/10.1038/s41598-021-86211-4, PMID 33767237.
-
Rothstein JD. Edaravone: a new drug approved for ALS. Cell. 2017 Nov 2;171(4):725. doi: 10.1016/j.cell.2017.10.011, PMID 29100067.
-
Bensimon G, Lacomblez L, Meininger V. A controlled trial of riluzole in amyotrophic lateral sclerosis. ALS/Riluzole study group. N Engl J Med. 1994 Mar 3;330(9):585-91, PMID 8302340.
-
Shefner J, Heiman Patterson T, Pioro EP, Wiedau‐Pazos M, Liu S, Zhang J, Agnese W, Apple S. Long‐term edaravone efficacy in amyotrophic lateral sclerosis: post‐hoc analyses of Study 19 (MCI186‐19). Muscle and Nerve. 2020 Feb;61(2):218-21. https://doi.org/10.1002/mus.26740, PMID 31621933.